Protein folding: the grand challenge of biology is still unsolved?
Where did the word ‘protein' come from and why? In 1838 Berzelius in his letter to Mulder discussing their results of elementary analysis of Albumin and Fibrin coined the term protein from the greek word ‘proteios’ which means primary or of primary importance, as he considered proteins where primary substance in animal nutrition.
Fundamentally proteins are biomolecules with a 3 dimensional shape. Proteins play a vital role in animal biology . Shortly speaking they are work doers of animal biology . I won't go into more details of the uses of proteins. But an important point to remember which we will keep coming up again and again in our discussion is that proteins are made from amino acids . These amino acids attach to each other to make a long linear chain known as polypeptide chain. This linear chain is also known as the primary structure of protein . Proteins need to get folded into a 3 dimensional shape for it to be biologically functional. Function of protein is only determined by its 3 dimensional shape. Determining protein structure experimentally is a costly approach approximately $100,000 per protein. Determining protein structure computationally is the only feasible approach to go forward. For proteins whose structures are determined experimentally earlier there is no issue but for new proteins whose structures are yet to be determined cost is an issue. We have to find an alternative to experiment . But how if we cannot determine protein shape experimentally then how can we even think of knowing proteins 3d shape and its function . If we can't do it experimentally then how can we do it computationally, what will be our inputs?
Yes it is true determining protein structure experimentally is expensive but determining sequence of amino acids in a protein is comparatively cheaper, approximately $1000 . It is known as whole genome sequencing. For a new protein we can easily know what the genome sequence of that protein is but it is hard for us to know its 3d structure. Knowing the genome sequence of a protein is of no use because only shape determines property of protein and not sequence. Then is genome sequence useless ?
The protein folding challenge
In 1950 Christiaan Anfinsen made a series of discoveries which showed that the information required to fold a polypeptide chain into a well defined and biologically functional conformation is strictly encoded in its amino acid sequence. After this biologists realized theoretically we only need an amino acid sequence to know a protein's 3 dimensional structure. We still to this date don’t know the physicochemical description of protein folding principle .In other words we don't know how protein fold from its primary structure to its final 3 dimensional form and this is protein folding challenge
Okay, now if we don't have an underlying principle neither it is economically feasible to determine protein structure experimentally then what to do now? There are many sequences for which we have experimentally found out 3 dimensional structure but we can learn nothing from patterns of those sequences so we started employing machines to learn those patterns . This process of machines learning patterns is known as machine learning
Machine learning in short (for those who have no idea)
Machine learning is a process through which we teach machines (computers) to learn from patterns in a given data. Machine learning can be categorized into three categories depending on the training process; supervised , unsupervised and reinforcement. Supervised learning is when we have input and expected output from that input and we train our machine so that it can learn to predict the right output for the purpose of this discussion. This is all that we will be needing . Machine learning consists of three parts: inputs, model and output . We give some input to the model (or computer algorithm) then the model gives us output , initially it will be bad at predicting and so we have a training process in which it learns from its mistake and at the end it learns to give the right prediction . To perform machine learning we need to have data (bigger the better) and there should be some correlation between our inputs and expected outputs. It is okay if we don't know the correlation , the machine will find it .
So by far what we have
Proteins 3 dimensional structure is strictly dictated by its amino acid sequence
We don't know underlying principle of protein folding so we cannot predict its structure from its sequence
It is expensive to find protein sequence experimentally so we are trying to find it computationally
Machine learning can be applied every where we have data and there is some correlation between inputs and outputs and from the first point we know there is some correlation
It is easy now we have data we can tell machines to learn on that data and the problem is solved . Eureka!
But no it is not the case . In theory the above sentence is true but in practicality it is not possible as of now. Simple reason is we just don't have enough data. The data we have currently is huge as per 2020 total residues in Protein Data Bank are 150423 but that is not enough for machines to learn .
So where do we go from here?
In the field of machine learning there are many such cases where we just don't have enough data. But there are many different techniques developed in the field of machine learning which can be applied that do not give perfect results but do give better than what we are having now or better than humans. Application of machine learning in the field of protein structure prediction is for sure one more step towards solving this problem.
So now let's explore application of machine learning in field of protein structure prediction
Broadly machine learning in protein structure prediction can be divided into 2 categories
Template based prediction : we use this approach for proteins for which we have homologous sequence or structure available in Protein Data Bank
De novo protein structure prediction : for orphan proteins we use this approach . Here we have to build protein structure from scratch
Inputs
In theory the amino acid sequence is all you need to predict the 3 dimensional structure of a protein , but in practice it is not possible because we are not having enough data so we use some additional information as input to our model . What is this additional information ? Different machine learning algorithms use different inputs. New state of the art models mostly use raw MSA’s as their input and some use templates too. What is a MSA and template?
Multiple sequence alignment (MSA) is a collection of all sequences that are similar to each other sequence wise or are evolutionarily related to each other. We do this in order to have more patterns to learn from. MSA is not perfect, it has its own flaws like many times it contains evolutionarily unrelated or irrelevant information . Initial attempts yielded mixed results but AlphaFold-2 achieved unprecedented efficiency using raw MSA . (This is discussed in detail why and how in the models section)
Position specific scoring matrix (PSSM) also known as position weight matrix(PWM), position specific weight matrix (PSWM). In PSSM we calculate observed frequencies on different amino acids at every residual position in a MSA. PSSM tells us which residue position are more highly conserved and their residue position
In some models MSA’s are also converted to Markov Random Field (MRF)
Templates are structures of homologous proteins or proteins that are evolutionarily related to our target protein. We use these structures as input to the network and predict the structure of our protein . Bellow this processes are described in details in machine learning models section
Different methods used for protein structure prediction
Template based modeling (TBM)
Template based modeling can be described in 4 steps
Finding homologous structure (templates related to the target protein sequence)
Aligning the target sequence to the template structure
Building structural frameworks by copying the aligned regions or by satisfying the spatial restrains from templates
Constructing the unaligned loop and adding side chain atoms
In a single procedure called threading (or fold recognition) we perform the first two steps as the correct template relies on accurate alignment. The last two steps are performed simultaneously as atoms of core and loop region are in close intact
Before neural networks
- Before neural networks, methods that relied on knowledge of physics were used . Rosetta is an example of such model
Beginning of machine learning in protein structure prediction
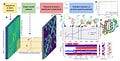
Initial machine learning algorithms used to take protein sequence and search for homologous families . This information was used to make MSA. This MSA was used to make MRF. These MRF were inputs to a type of neural network architecture called Convolutional neural networks (CNN) . In Early 2015 2 dimensional MRF’s were treated like images so that we can utilize techniques developed in computer vision . They returned binary contact map as their output
RaptorX
RaptorX introduced Resnet to protein structure prediction . with the help of Resnet CNN started to improve contact map quality drastically . ResNets helped in training very deep CNN with hundreds of layers enabling wider receptive fields and greater expressive power . As a result contact maps were more accurate as compared to older methods . RaptorX changed everyone’s perception of using deep learning techniques in protein structure prediction and everyone was motivated to explore more into application of deep learning in protein structure prediction problems . RaptorX was the architecture of choice for all models before 2020. All models after RaptorX were made on RaptorX architecture with some changes in output
AlphaFold 1
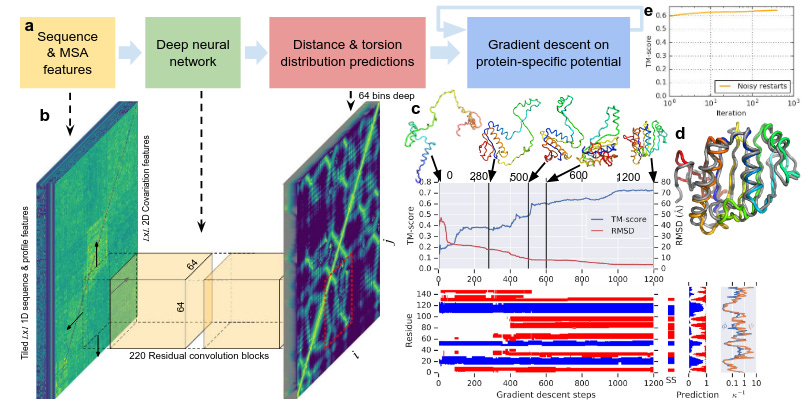
AlphaFold 1 was a major success when it was released it performed better than many different algorithms available at that time . AlphaFold1 used Resnet’s for predicting protein structure . AlphaFold1 predicted the probability distribution of distances between any two residues . It outputted distograms and orientograms
Despite these major improvement we were not still at a place where we can say protein folding problem is “solved”
AlphaFold 2
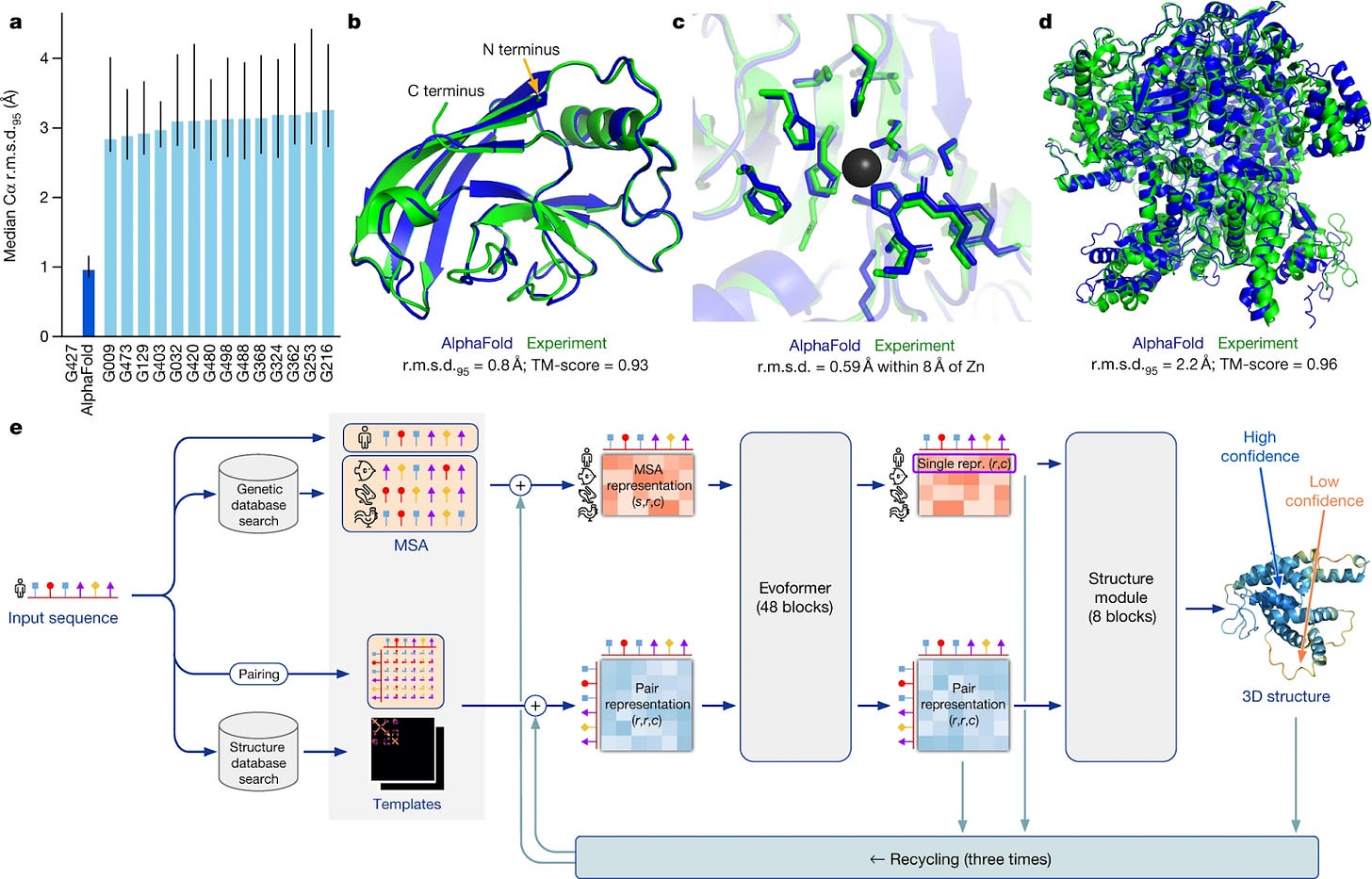
Alphafold2 was using a transformer over CNN or ResNets. Instead of using only raw MSA as input AlphaFold2 used MSA and templates as input to their system and one more major change in AlphaFold2 was it started to output 3d structure consistent in euclidean space . For structures whose evolutionary information is available AlphaFold2 predicted highly accurate results. But for structures with no evolutionary information AlphaFold2 could not perform well
With the success of AlphaFold2 we can say that for proteins whose evolutionary information is available we can predict its 3D structure
If you have any feedback/suggestions email me at patilabhishekb25902@gmail.com